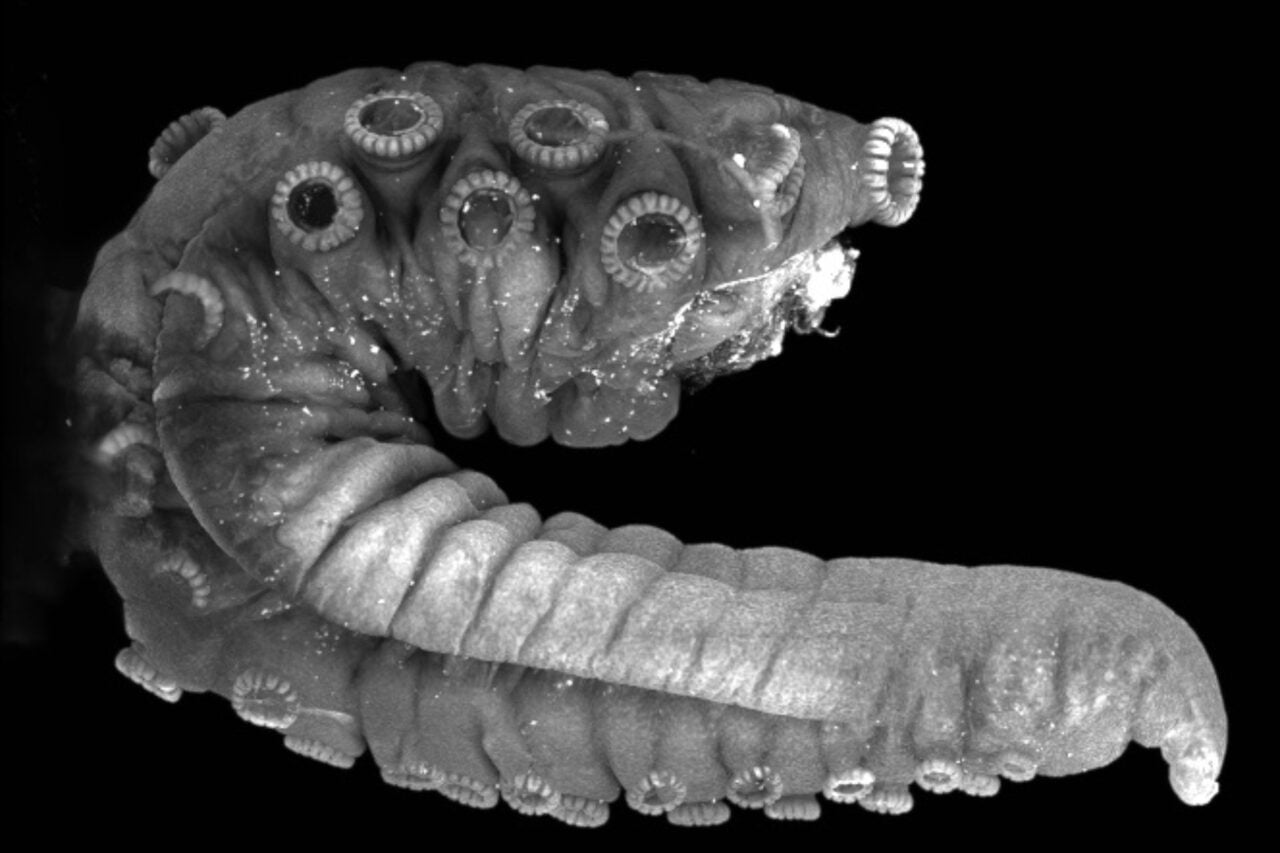
By building a gigantic petri dish, researchers from Harvard Medical School and Technion-Israel Institute of Technology have produced a jaw-dropping visualization showing bacteria as it mutates to become resistant to drugs.
The new study, published today in Science, is the first large-scale demonstration showing how bacteria react to ever-increasing doses of antibiotics, and how these relentless microbes exploit Darwinian selection to adapt to—and even thrive within—the very medicines meant to kill them.
“What surprised me most about it was that we could actually see evolution happening in front of us,” co-author Michael Baym, a postdoc at the Kishony lab at Harvard Medical School, told Gizmodo. “Here were the abstract diagrams we’d been drawing for years, come to life.”
https://gizmodo.com/antibiotic-resistant-superbugs-could-kill-10-million-pe-1777549058
Each year, around 700,000 people die around the world from untreatable bacterial infections, and antibiotic-resistant superbugs could kill upwards of 10 million people each year by the mid-21st century. Just today, the UN announced a high-level meeting to discuss possible strategies and countermeasures.
Baym worked with Roy Kishony of Technion-Israel Institute of Technology and Harvard Medical School on the experiment. They call their giant petri dish the Microbial Evolution and Growth Arena, or MEGA for short. It’s a rectangular platform, two feet wide and four feet long, filled with a gelatinous substance known as agar, a seaweed-derived substance that’s commonly used to facilitate microbial growth. Using the MEGA-plate, the researchers were able to watch antibiotic resistance develop in Escherichia coli.
They divided the MEGA-plate into several sections, each of which was saturated with varying doses of antibiotics. The ends of the platform contained no antibiotics, allowing the bacteria to thrive; these areas represented the starting line. But the adjacent inner sections contained a small amount of antibiotic—just enough to kill the E. coli. Moving inward, each subsequent section of the MEGA plate was treated with a ten-fold increase in the dose of antibiotics. At the very core of the dish, there was 1,000 times as much antibiotic compared to the areas with the lowest dose.
For the next two weeks, the researchers watched—and filmed—as the bacteria died, survived, and adapted to the increasingly poisonous conditions located at the borders of their immediate perimeters. The resulting timelapse video literally shows Darwinian processes at work—a process that would normally remain invisible to the human eye.
As the two-week experiment progressed, the bacteria spread until they reached a potent concentration of antibiotics beyond which they could not grow. That is, until mutants—armed with the specific set of traits required to fight off the poison—finally emerged. This often didn’t take long. At each concentration level, a small segment of bacteria adapted to the hostile conditions, the result of successive accumulated genetic changes.
Once settled in the new section of the MEGA-plate, these tiny populations of antibacterial-resistant mutants were able to grow. When they reached the next section of the platform, the pattern repeated itself. The descendents of this initial group of mutants were able to move to areas filled with higher concentrations of antibiotics. Eventually, multiple lineages of mutants competed for the same space, with winning strains moving on to areas with higher drug doses.
By the eleventh day, the bacteria had migrated all the way to the highest drug concentration in the center. These hardy mutants were capable of surviving an antibiotic known as trimethoprim at a dose 1,000 times greater than the one that killed their ancestors. And some bacteria acquired a 100,000-fold ability to fend off ciprofloxacin, another common antibiotic.
“We were able to evolve over a thousand-fold wild-type resistance to trimethoprim in 11 days— that’s very nearly the saturation limit of the drug,” said Baym. “Put simply, there was no way to dissolve enough drug to kill these bacteria.” Importantly, all bacterial mutants were contained and all materials decontaminated after use.
Observations showed that initial mutations led to slower growth. That suggests bacteria aren’t capable of growing at optimal speeds while in the midst of developing adaptations. But once they stumble upon a fortuitous immunity, it’s all systems go, with growth proceeding at normal rates.
Also, the fittest mutants weren’t always the fastest growers. The most successful bacteria remained behind while the weaker strains were forced to deal with the intense drug doses at the front lines.
“Thanks to the the bacteria needing to migrate to survive, we saw a surprising dynamic by which the strongest weren’t necessarily winning, rather those that were good enough and close enough to the new area would beat out nominally superior mutants just by being faster,” Baym said. “Nevertheless, in every case we saw that this successive accumulation of mutations was able to evolve extremely high levels of antibiotic resistance in a relatively short time.”
Looking ahead, the researchers would like to use the the MEGA-plate to predict the future evolutionary potential of specific pathogens. Armed with this knowledge, future clinicians will be able to tell which antibiotic a pathogen is resistant to, and how it might evolve resistance if certain antibiotics are used.
[Science]