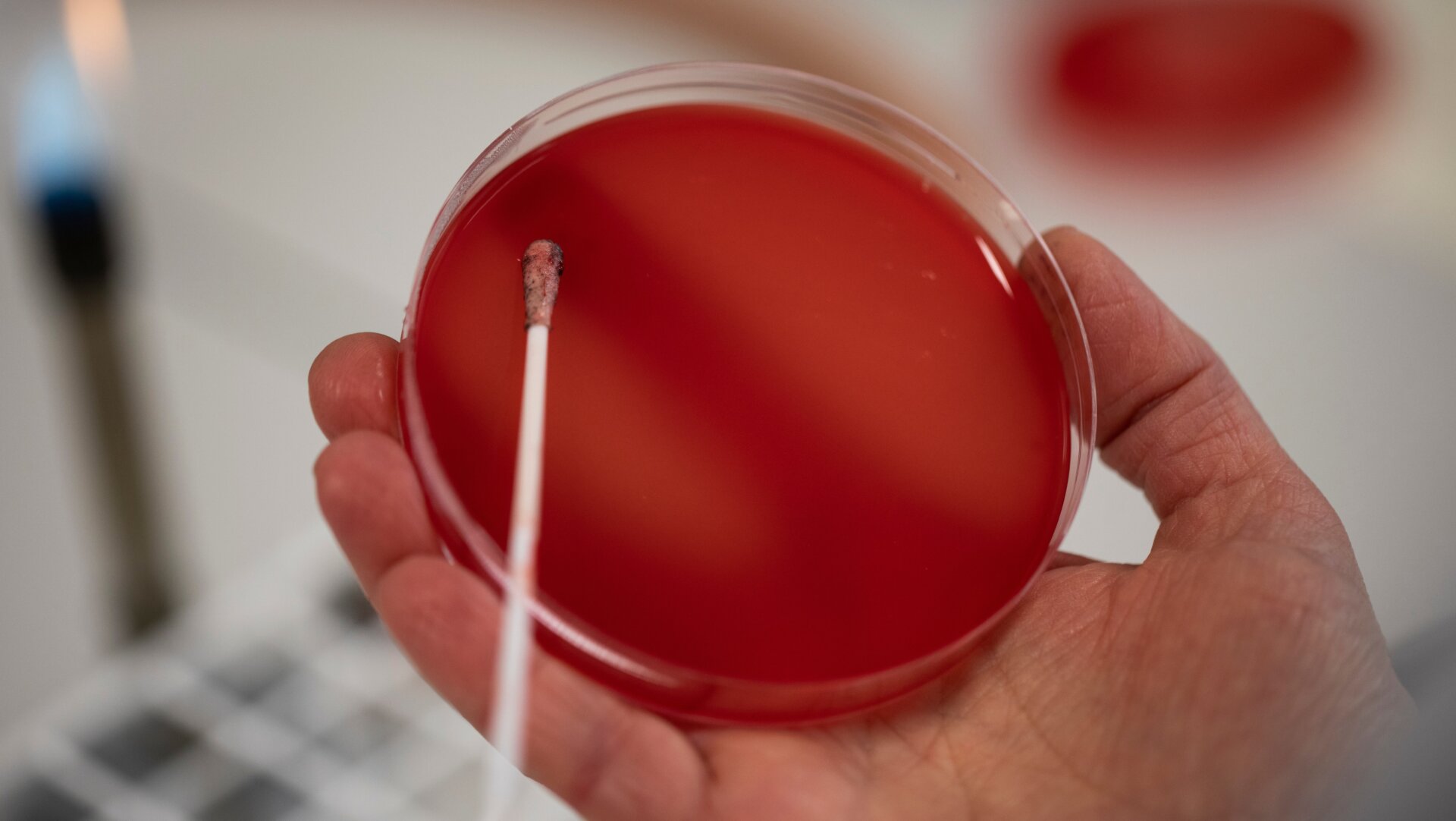
Here’s a little morsel of horror to carry you through the weekend. Scientists at North Carolina State University say they’ve discovered disease-causing Salmonella bacteria in the U.S. that can resist one of the few remaining antibiotics available for certain infections. What’s worse, the mutation responsible for this newfound resistance can easily be passed onto other bacteria, raising the risk of some infections eventually becoming untreatable with antibiotics.
The last-resort antibiotic in question is a 50-year-old drug called colistin. In its heyday, colistin was hardly the most safe and effective treatment, and it eventually fell out of favor with doctors because it could cause serious kidney damage. But it has reemerged as a last line of defense recently because it can still kill gram-negative bacteria resistant to multiple antibiotics. These bacteria include commonly encountered bugs like Escherichia coli as well as hospital-associated infections caused by Pseudomonas aeruginosa and Klebsiella pneumoniae.
So far, there haven’t been any signs of widespread resistance to colistin among these specific superbugs. But scientists have started to spot genetic mutations that allow bacteria to resist colistin in other bacteria—on bits of free-moving DNA called plasmids. After the initial discovery of a mobilized colistin resistance (MCR) gene was published in 2015, at least two but possibly as many as eight other MCR variants have been found elsewhere.
https://gizmodo.com/fecal-transplant-patient-killed-by-superbug-traced-to-d-1835521476
What makes this so dangerous is that plasmids can move between bacteria. So it’s possible these MCR genes can find their way to a strain of E. coli resistant to all other drugs, for example. And if that strain infects and seriously sickens a person, there might be no conceivable way for doctors to treat it. If this event happens again and again—and barring the discovery of a new or old drug that can step in where colistin failed—we’d be left with a reality where some infections, especially those in hospitals or other areas where superbugs are prevalent, are simply untreatable.
It’s this nightmare scenario that’s compelled doctors, including the authors behind this latest study, to start aggressively looking for resistance genes, particularly MCR, in the wild.
According to the study, the find was made in an 18-year-old female patient who came down with food poisoning, namely diarrhea, and visited a doctor in 2014. After testing, it was suspected that she had become sick from a strain of multidrug-resistant Salmonella. When the team tested her sample, though, they also found a variant of MCR, MCR3-1, on a plasmid.
Their work was published earlier this June in the Journal of Medical Microbiology.
Colistin isn’t typically used to treat Salmonella, though it is related to E. coli and other gram negative bacteria. And the woman’s infection was still treatable with other antibiotics (due to medical confidentiality, though, her ultimate fate isn’t known). But the discovery of this particular combination of Salmonella bacteria and MCR 3.1—the first known instance, according to the researchers—is ominous. Two weeks before his doctor’s visit, he had been traveling in China. The country’s rampant use of colistin as a farm antibiotic, which only recently ended, is thought to have sped up the emergence of MCR. But as this case shows, MCR has long since gone international, having been spotted in more than two dozen countries in both people and animals.
These MCR cases are not untreatable (or even always disease-causing) for the time being. But the more often they occur, the greater the odds that truly pan-resistant and serious infections will become the new normal. Given that this case actually happened in 2014, it’s also possible that the overall situation has only gotten worse since.
All of which is to say: Have a good weekend!
Update: This article previously referred to the patient as male. Senior study author Siddhartha Thakur told Gizmodo that while the research paper described the patient as male, she was actually female, and the study will be corrected in the near future.